Autism Spectrum Disorders (ASDs) are neurodevelopmental disorders (NDDs) in which children display differences in social interaction/communication and repetitive stereotyped behaviors along with variable associated features. Cul3, a gene linked to ASD, encodes CUL3 (CULLIN-3), a protein that serves as a key component of a ubiquitin ligase complex with unclear function in neurons. Cul3 homozygous deletion in mice is embryonic lethal; thus, we examine the role of Cul3 deletion in early synapse development and neuronal morphology in hippocampal primary neuronal cultures. Homozygous deletion of Cul3 significantly decreased dendritic complexity and dendritic length, as well as axon formation. Synaptic spine density significantly increased, mainly in thin and stubby spines along with decreased average spine volume in Cul3 knockouts. Both heterozygous and homozygous knockout of Cul3 caused significant reductions in the density and colocalization of gephyrin/vGAT puncta, providing evidence of decreased inhibitory synapse number, while excitatory synaptic puncta vGulT1/PSD95 density remained unchanged. Based on previous studies implicating elevated caspase-3 after Cul3 deletion, we demonstrated increased caspase-3 in our neuronal cultures and decreased neuronal cell viability. We then examined the efficacy of the caspase-3 inhibitor Z-DEVD-FMK to rescue the decrease in neuronal cell viability, demonstrating reversal of the cell viability phenotype with caspase-3 inhibition. Studies have also implicated caspase-3 in neuronal morphological changes. We found that caspase-3 inhibition largely reversed the dendrite, axon, and spine morphological changes along with the inhibitory synaptic puncta changes. Overall, these data provide additional evidence that Cul3 regulates the formation or maintenance of cell morphology, GABAergic synaptic puncta, and neuronal viability in developing hippocampal neurons in culture.
Copyright © 2024 Xia, Singh, Wang, Xuan, Singer and Powell.
Product Citations: 9
In Frontiers in Cellular Neuroscience on 28 May 2024 by Xia, Q. Q., Singh, A., et al.
-
WB
-
Mus musculus (House mouse)
-
Neuroscience
Effects of heterozygous deletion of autism-related gene Cullin-3 in mice.
In PLoS ONE on 10 July 2023 by Xia, Q. Q., Walker, A. K., et al.
Autism Spectrum Disorder (ASD) is a developmental disorder in which children display repetitive behavior, restricted range of interests, and atypical social interaction and communication. CUL3, coding for a Cullin family scaffold protein mediating assembly of ubiquitin ligase complexes through BTB domain substrate-recruiting adaptors, has been identified as a high-risk gene for autism. Although complete knockout of Cul3 results in embryonic lethality, Cul3 heterozygous mice have reduced CUL3 protein, demonstrate comparable body weight, and display minimal behavioral differences including decreased spatial object recognition memory. In measures of reciprocal social interaction, Cul3 heterozygous mice behaved similarly to their wild-type littermates. In area CA1 of hippocampus, reduction of Cul3 significantly increased mEPSC frequency but not amplitude nor baseline evoked synaptic transmission or paired-pulse ratio. Sholl and spine analysis data suggest there is a small yet significant difference in CA1 pyramidal neuron dendritic branching and stubby spine density. Unbiased proteomic analysis of Cul3 heterozygous brain tissue revealed dysregulation of various cytoskeletal organization proteins, among others. Overall, our results suggest that Cul3 heterozygous deletion impairs spatial object recognition memory, alters cytoskeletal organization proteins, but does not cause major hippocampal neuronal morphology, functional, or behavioral abnormalities in adult global Cul3 heterozygous mice.
Copyright: © 2023 Xia et al. This is an open access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
-
WB
-
Neuroscience
In IScience on 21 April 2023 by Dubiel, D., Wang, J., et al.
The COP9 signalosome (CSN) and cullin-RING ubiquitin ligases (CRLs) form latent CSN-CRL complexes detectable in cells. We demonstrate that the CSN variants CSNCSN7A and CSNCSN7B preferentially bind to CRL3 or CRL4A and CRL4B, respectively. Interestingly, the interacting protein ubiquitin-specific protease 15 exclusively binds to latent CSNCSN7A-CRL3, while p27KIP attaches to latent CSNCSN7B-CRL4A complex. Inhibition of deneddylation by CSN5i-3 or neddylation by MLN4924 do not impede the formation of latent complexes. Latent CSNCSN7A-CRL3 and latent CSNCSN7B-CRL4A/B particles are essential for specific cellular functions. We found that curcumin-induced cell death requires latent CSNCSN7B-CRL4A. Knockout of CSN7B in HeLa cells leads to resistance against curcumin. Remarkably, the small GTPase RAB18 recruits latent CSNCSN7A-CRL3 complex to lipid droplets (LDs), where CRL3 is activated by neddylation, an essential event for LD formation during adipogenesis. Knockdown of CSN7A or RAB18 or destabilization of latent complexes by cutting off CSN7A C-terminal 201-275 amino acids blocks adipogenesis.
© 2023 The Author(s).
-
Homo sapiens (Human)
In Proceedings of the National Academy of Sciences of the United States of America on 23 November 2021 by Lee, J., Lim, C., et al.
Circadian transcriptional timekeepers in pacemaker neurons drive profound daily rhythms in sleep and wake. Here we reveal a molecular pathway that links core transcriptional oscillators to neuronal and behavioral rhythms. Using two independent genetic screens, we identified mutants of Transport and Golgi organization 10 (Tango10) with poor behavioral rhythmicity. Tango10 expression in pacemaker neurons expressing the neuropeptide PIGMENT-DISPERSING FACTOR (PDF) is required for robust rhythms. Loss of Tango10 results in elevated PDF accumulation in nerve terminals even in mutants lacking a functional core clock. TANGO10 protein itself is rhythmically expressed in PDF terminals. Mass spectrometry of TANGO10 complexes reveals interactions with the E3 ubiquitin ligase CULLIN 3 (CUL3). CUL3 depletion phenocopies Tango10 mutant effects on PDF even in the absence of the core clock gene timeless Patch clamp electrophysiology in Tango10 mutant neurons demonstrates elevated spontaneous firing potentially due to reduced voltage-gated Shaker-like potassium currents. We propose that Tango10/Cul3 transduces molecular oscillations from the core clock to neuropeptide release important for behavioral rhythms.
Copyright © 2021 the Author(s). Published by PNAS.
-
WB
RhoBTB1 interacts with ROCKs and inhibits invasion.
In Biochemical Journal on 13 September 2019 by Haga, R. B., Garg, R., et al.
RhoBTB1 is an atypical Rho GTPase with two BTB domains in addition to its Rho domain. Although most Rho GTPases regulate actin cytoskeletal dynamics, RhoBTB1 is not known to affect cell shape or motility. We report that RhoBTB1 depletion increases prostate cancer cell invasion and induces elongation in Matrigel, a phenotype similar to that induced by depletion of ROCK1 and ROCK2. We demonstrate that RhoBTB1 associates with ROCK1 and ROCK2 and its association with ROCK1 is via its Rho domain. The Rho domain binds to the coiled-coil region of ROCK1 close to its kinase domain. We identify two amino acids within the Rho domain that alter RhoBTB1 association with ROCK1. RhoBTB1 is a substrate for ROCK1, and mutation of putative phosphorylation sites reduces its association with Cullin3, a scaffold for ubiquitin ligases. We propose that RhoBTB1 suppresses cancer cell invasion through interacting with ROCKs, which in turn regulate its association with Cullin3. Via Cullin3, RhoBTB1 has the potential to affect protein degradation.
© 2019 The Author(s).
-
Homo sapiens (Human)
-
Biochemistry and Molecular biology
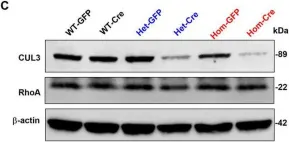
In Front Cell Neurosci on 28 May 2024 by Xia, Q. Q., Singh, A., et al.
Fig.1.C

-
WB
-
Mus musculus (House mouse)
Collected and cropped from Front Cell Neurosci by CiteAb, provided under a CC-BY license
Image 1 of 4
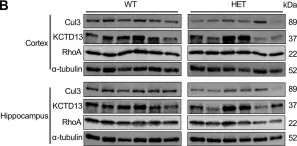
In PLoS One on 10 July 2023 by Xia, Q. Q., Walker, A. K., et al.
Fig.1.B

-
WB
-
Collected and cropped from PLoS One by CiteAb, provided under a CC-BY license
Image 1 of 4
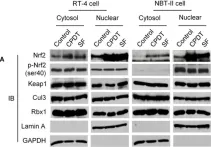
In PLoS One on 5 May 2012 by Li, Y., Paonessa, J. D., et al.
Fig.3.A

-
WB
-
Collected and cropped from PLoS One by CiteAb, provided under a CC-BY license
Image 1 of 4
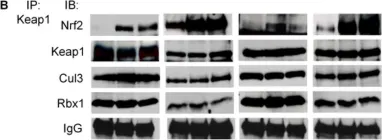
In PLoS One on 5 May 2012 by Li, Y., Paonessa, J. D., et al.
Fig.3.B

-
WB
-
Collected and cropped from PLoS One by CiteAb, provided under a CC-BY license
Image 1 of 4