Metabolic demands fluctuate rhythmically and rely on coordination between the circadian clock and nutrient-sensing signalling pathways, yet mechanisms of their interaction remain not fully understood. Surprisingly, we find that class 3 phosphatidylinositol-3-kinase (PI3K), known best for its essential role as a lipid kinase in endocytosis and lysosomal degradation by autophagy, has an overlooked nuclear function in gene transcription as a coactivator of the heterodimeric transcription factor and circadian driver Bmal1-Clock. Canonical pro-catabolic functions of class 3 PI3K in trafficking rely on the indispensable complex between the lipid kinase Vps34 and regulatory subunit Vps15. We demonstrate that although both subunits of class 3 PI3K interact with RNA polymerase II and co-localize with active transcription sites, exclusive loss of Vps15 in cells blunts the transcriptional activity of Bmal1-Clock. Thus, we establish non-redundancy between nuclear Vps34 and Vps15, reflected by the persistent nuclear pool of Vps15 in Vps34-depleted cells and the ability of Vps15 to coactivate Bmal1-Clock independently of its complex with Vps34. In physiology we find that Vps15 is required for metabolic rhythmicity in liver and, unexpectedly, it promotes pro-anabolic de novo purine nucleotide synthesis. We show that Vps15 activates the transcription of Ppat, a key enzyme for the production of inosine monophosphate, a central metabolic intermediate for purine synthesis. Finally, we demonstrate that in fasting, which represses clock transcriptional activity, Vps15 levels are decreased on the promoters of Bmal1 targets, Nr1d1 and Ppat. Our findings open avenues for establishing the complexity for nuclear class 3 PI3K signalling for temporal regulation of energy homeostasis.
© 2023. The Author(s).
Applications
Reactivity
Research Area
Class 3 PI3K coactivates the circadian clock to promote rhythmic de novo purine synthesis.
In Nature Cell Biology on 1 July 2023 by Alkhoury, C., Henneman, N. F., et al.
-
ChIP
-
IP
-
IF
-
WB
-
Homo sapiens (Human)
-
Mus musculus (House mouse)
-
Cell Biology
-
Genetics
In Autophagy on 1 August 2019 by Birgisdottir, Å. B., Mouilleron, S., et al.
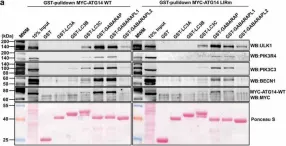
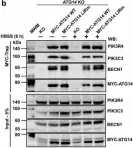
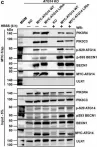
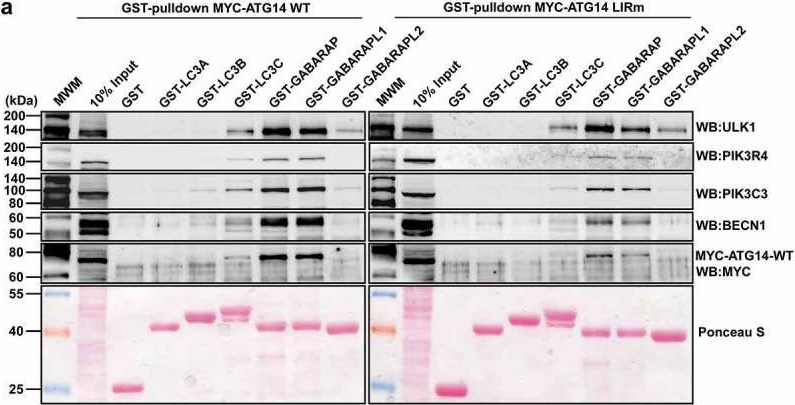
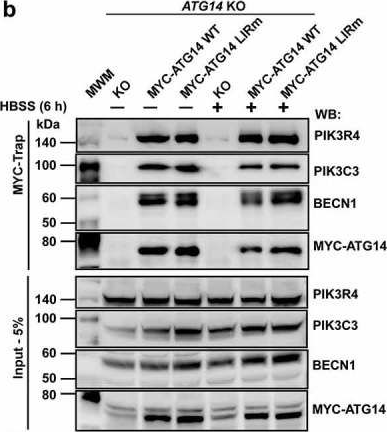
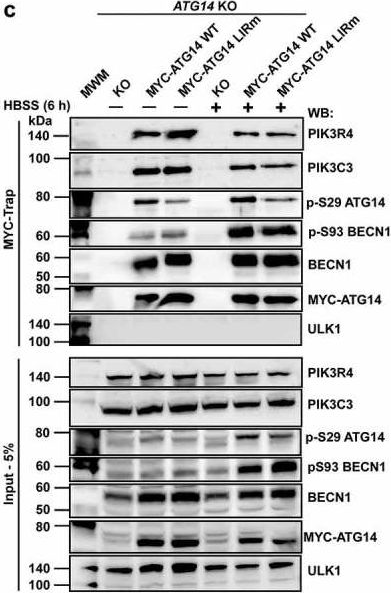
Autophagosome formation depends on a carefully orchestrated interplay between membrane-associated protein complexes. Initiation of macroautophagy/autophagy is mediated by the ULK1 (unc-51 like autophagy activating kinase 1) protein kinase complex and the autophagy-specific class III phosphatidylinositol 3-kinase complex I (PtdIns3K-C1). The latter contains PIK3C3/VPS34, PIK3R4/VPS15, BECN1/Beclin 1 and ATG14 and phosphorylates phosphatidylinositol to generate phosphatidylinositol 3-phosphate (PtdIns3P). Here, we show that PIK3C3, BECN1 and ATG14 contain functional LIR motifs and interact with the Atg8-family proteins with a preference for GABARAP and GABARAPL1. High resolution crystal structures of the functional LIR motifs of these core components of PtdIns3K-C1were obtained. Variation in hydrophobic pocket 2 (HP2) may explain the specificity for the GABARAP family. Mutation of the LIR motif in ATG14 did not prevent formation of the PtdIns3K-C1 complex, but blocked colocalization with MAP1LC3B/LC3B and impaired mitophagy. The ULK-mediated phosphorylation of S29 in ATG14 was strongly dependent on a functional LIR motif in ATG14. GABARAP-preferring LIR motifs in PIK3C3, BECN1 and ATG14 may, via coincidence detection, contribute to scaffolding of PtdIns3K-C1 on membranes for efficient autophagosome formation. Abbreviations: ATG: autophagy-related; BafA1: bafilomycin A1; GABARAP: GABA type A receptor-associated protein; GABARAPL1: GABA type A receptor associated protein like 1; GFP: enhanced green fluorescent protein; KO: knockout; LDS: LIR docking site; LIR: LC3-interacting region; MAP1LC3/LC3: microtubule associated protein 1 light chain 3; PIK3C3: phosphatidylinositol 3-kinase catalytic subunit type 3; PIK3R4: phosphoinositide-3-kinase regulatory subunit 4; PtdIns3K: phosphatidylinositol 3-kinase; PtdIns3P: phosphatidylinositol-3-phosphate; SQSTM1/p62: sequestosome 1; VPS: Vacuolar protein sorting; ULK: unc-51 like autophagy activating kinase.
-
WB
-
Homo sapiens (Human)
-
Cell Biology
In Nature Communications on 18 January 2018 by Miranda, A. M., Lasiecka, Z. M., et al.
Defects in endolysosomal and autophagic functions are increasingly viewed as key pathological features of neurodegenerative disorders. A master regulator of these functions is phosphatidylinositol-3-phosphate (PI3P), a phospholipid synthesized primarily by class III PI 3-kinase Vps34. Here we report that disruption of neuronal Vps34 function in vitro and in vivo impairs autophagy, lysosomal degradation as well as lipid metabolism, causing endolysosomal membrane damage. PI3P deficiency also promotes secretion of unique exosomes enriched for undigested lysosomal substrates, including amyloid precursor protein C-terminal fragments (APP-CTFs), specific sphingolipids, and the phospholipid bis(monoacylglycero)phosphate (BMP), which normally resides in the internal vesicles of endolysosomes. Secretion of these exosomes requires neutral sphingomyelinase 2 and sphingolipid synthesis. Our results reveal a homeostatic response counteracting lysosomal dysfunction via secretion of atypical exosomes eliminating lysosomal waste and define exosomal APP-CTFs and BMP as candidate biomarkers for endolysosomal dysfunction associated with neurodegenerative disorders.
-
WB
-
Mus musculus (House mouse)
-
Cell Biology
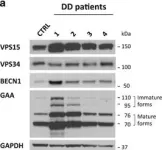
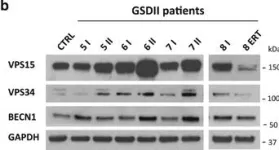
Autophagy dysregulation in Danon disease.
In Cell Death & Disease on 19 January 2017 by Nascimbeni, A. C., Fanin, M., et al.
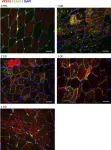
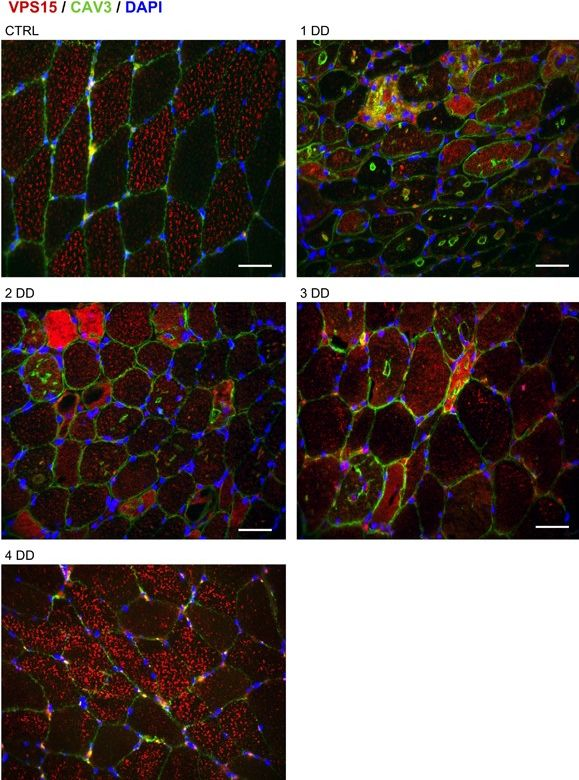
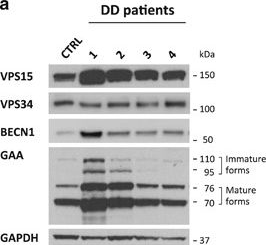
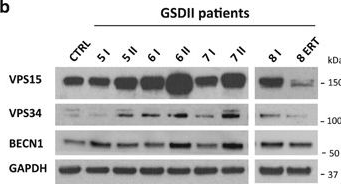
The autophagy-lysosome system is critical for muscle homeostasis and defects in lysosomal function result in a number of inherited muscle diseases, generally referred to as autophagic vacuolar myopathies (AVMs). Among them, Danon Disease (DD) and glycogen storage disease type II (GSDII) are due to primary lysosomal protein defects. DD is characterized by mutations in the lysosome-associated membrane protein 2 (LAMP2) gene. The DD mouse model suggests that inefficient lysosome biogenesis/maturation and impairment of autophagosome-lysosome fusion contribute to the pathogenesis of muscle wasting. To define the role of autophagy in human disease, we analyzed the muscle biopsies of DD patients and monitored autophagy and several autophagy regulators like transcription factor EB (TFEB), a master player in lysosomal biogenesis, and vacuolar protein sorting 15 (VPS15), a critical factor for autophagosome and endosome biogenesis and trafficking. Furthermore, to clarify whether the mechanisms involved are shared by other AVMs, we extended our mechanistic study to a group of adult GSDII patients. Our data show that, similar to GSDII, DD patients display an autophagy block that correlates with the severity of the disease. Both DD and GSDII show accumulation and altered localization of VPS15 in autophagy-incompetent fibers. However, TFEB displays a different pattern between these two lysosomal storage diseases. Although in DD TFEB and downstream targets are activated, in GSDII patients TFEB is inhibited. These findings suggest that these regulatory factors may have an active role in the pathogenesis of these diseases. Therapeutic approaches targeted to normalize these factors and restore the autophagic flux in these patients should therefore be considered.
-
IHC-IF
-
WB
-
IF
-
Homo sapiens (Human)
-
Cell Biology
In Cellular Signalling on 1 June 2014 by Hirsch, D. S., Shen, Y., et al.
The class III phosphatidylinositol 3-kinase, VPS34, phosphorylates the D3 hydroxyl of inositol generating phosphatidylinositol 3-phosphate (ptdins(3)p). Initial studies suggested that ptdins(3)p solely functioned as a component of vesicular and endosomal membranes and that VPS34 did not function in signal transduction. However, VPS34 has recently been shown to be required for insulin-mediated activation of S6 kinase 1 (S6K1). Whether VPS34 activity is directly regulated by insulin is unclear. It is also not known whether VPS34 activity can be spatially restricted in response to extracellular stimuli. Data presented here demonstrate that in response to insulin, VPS34 is activated and translocated to lamellipodia where it produces ptdins(3)p. The localized production of ptdins(3)p is dependent on Src phosphorylation of VPS34. In cells expressing VPS34 with mutations at Y231 or Y310, which are Src-phosphorylation sites, insulin-stimulated VPS34 translocation to the plasma membrane and lamellipodia formation are blocked. mTOR also colocalizes with VPS34 and ptdins(3)p at lamellipodia following insulin-stimulation. In cells expressing the VPS34-Y231F mutant, which blocks lamellipodia formation, mTOR localization at the plasma membrane and insulin-mediated S6K1 activation are reduced. This suggests that mTOR localization at lamellipodia is important for full activation of S6K1 induced by insulin. These data demonstrate that insulin can spatially regulate VPS34 activity through Src-mediated tyrosine phosphorylation and that this membrane localized activity contributes to lamellipodia formation and activation of mTOR/S6K1signaling.
Published by Elsevier Inc.
-
IF
-
WB
-
Mus musculus (House mouse)
-
Chlorocebus sabaeus (Green monkey)
-
Cell Biology
-
Endocrinology and Physiology
In Autophagy on 1 August 2019 by Birgisdottir, Å. B., Mouilleron, S., et al.
Fig.7.A
-
WB
-
Homo sapiens (Human)
Collected and cropped from Autophagy by CiteAb, provided under a CC-BY license
Image 1 of 6
In Autophagy on 1 August 2019 by Birgisdottir, Å. B., Mouilleron, S., et al.
Fig.7.B
-
WB
-
Homo sapiens (Human)
Collected and cropped from Autophagy by CiteAb, provided under a CC-BY license
Image 1 of 6
In Autophagy on 1 August 2019 by Birgisdottir, Å. B., Mouilleron, S., et al.
Fig.7.C
-
WB
-
Homo sapiens (Human)
Collected and cropped from Autophagy by CiteAb, provided under a CC-BY license
Image 1 of 6
In Cell Death Dis on 19 January 2017 by Nascimbeni, A. C., Fanin, M., et al.
Fig.4.A
-
IHC-IF
-
Homo sapiens (Human)
Collected and cropped from Cell Death & Disease by CiteAb, provided under a CC-BY license
Image 1 of 6
In Cell Death Dis on 19 January 2017 by Nascimbeni, A. C., Fanin, M., et al.
Fig.3.A
-
WB
-
Homo sapiens (Human)
Collected and cropped from Cell Death & Disease by CiteAb, provided under a CC-BY license
Image 1 of 6
In Cell Death Dis on 19 January 2017 by Nascimbeni, A. C., Fanin, M., et al.
Fig.3.B
-
WB
-
Collected and cropped from Cell Death & Disease by CiteAb, provided under a CC-BY license
Image 1 of 6