Tumor cell plasticity drives metastasis and therapy resistance, yet its regulation by oncoprotein dosage dynamics remains poorly understood. In Ewing sarcoma (EwS), variations in EWS::FLI1 (EF) fusion oncoprotein activity have been associated with epithelial-mesenchymal plasticity (EMP). Using degron technology, we precisely modulated endogenous EF in EwS cells and linked phenotypic states to distinct oncoprotein dosages. Strikingly, modest EF depletion promoted a pro-metastatic phenotype that diminished upon near-complete EF loss, revealing a paradoxical effect of submaximal EF inhibition. Nascent RNA-sequencing uncovered distinct gene clusters with heterogenous transcriptional responses to graded EF loss. Genes most sensitive to subtle EF depletion harbored GGAA microsatellites within EF-bound enhancers, while chromatin profiling uncovered candidate cofactors regulating EF-repressed EMP programs. Transient EF depletion followed by rapid restoration, modelling oncoprotein fluctuations, caused persistent dysregulation of genes functionally linked to enhanced extravasation and metastatic burden in preclinical models. This study highlights the therapeutic challenge of incomplete EF elimination, serving a paradigm in which oncoprotein dosage dynamics act as non-genetic drivers of disease progression and reveal novel vulnerabilities of advanced disease.
© 2025. The Author(s).
Modelling EWS::FLI1 protein fluctuations reveal determinants of tumor plasticity in Ewing sarcoma.
In EMBO Molecular Medicine on 1 February 2026 by Suresh, V., Hafemeister, C., et al.
Kinetic measurement of gene-specific RNA polymerase II transcription elongation rates.
In Genome Research on 3 November 2025 by Liu, H. & Gregersen, L. H.
Transcription is regulated at multiple levels, including initiation, elongation, and termination. Whereas much research has focused on the initiation of transcription, regulation of elongation plays an important role not only in transcription dynamics but also in cotranscriptional RNA processing and genome stability. Despite advances in high-throughput approaches for global quantification of RNA polymerase II (RNAPII) speed, RNAPII elongation rate studies have been limited to a relatively small number of long genes or to velocity estimates inferred indirectly from RNAPII occupancy data. Here, we present DRB/TTchem-seq2, a modified version of the DRB/TTchem-seq method, to directly measure gene-specific elongation rates of more than 3000 genes. By combining short time point sampling after synchronized RNAPII release into the gene body and a new computational framework to track the distance traveled by RNAPII, we greatly increase the number of genes for which it is possible to obtain elongation rates. Our direct RNAPII elongation rate quantification reveals that elongation rates vary not only among genes but also within genes. Additionally, we describe how specific histone modifications and elongation factor occupancy correlate with subclasses of genes based on their elongation rates. Together, we present a robust and powerful method for RNAPII transcription elongation rate measurement.
© 2025 Liu and Gregersen; Published by Cold Spring Harbor Laboratory Press.
In Nucleic Acids Research on 20 June 2025 by Soderholm, A., Vunjak, M., et al.
Type II interferon (IFNγ) signaling is essential for innate immunity and critical for effective immunological checkpoint blockade in cancer immunotherapy. Genetic screen identification of post-transcriptional regulators of this pathway has been challenging since such factors are often essential for cell viability. Here, we utilize our inducible CRISPR/Cas9 approach to screen for key post-transcriptional regulators of IFNγ signaling, and in this way, we identify ERH and the ERH-associated splicing and RNA export factors MAGOH, SRSF1, and ALYREF. Loss of these factors impairs post-transcriptional mRNA maturation of JAK2, a crucial kinase for IFNγ signaling, resulting in abrogated JAK2 protein levels and diminished IFNγ signaling. Further analysis highlights a critical role for ERH in preventing intron retention in AU-rich regions in specific transcripts, such as JAK2. This regulation is markedly different from previously described retention of GC-rich introns. Overall, these findings reveal that post-transcriptional JAK2 processing is a critical rate-limiting step for the IFNγ-driven innate immune response.
© The Author(s) 2025. Published by Oxford University Press on behalf of Nucleic Acids Research.
Preprint on BioRxiv : the Preprint Server for Biology on 8 April 2025 by Suresh, V., Hafemeister, C., et al.
ABSTRACT The mechanisms underlying tumor cell plasticity driving drug resistance and disease progression remain poorly understood. In Ewing sarcoma (EwS), variations in EWS::FLI1 (EF) activity have been associated with epithelial-mesenchymal plasticity (EMP). Using degron technology, we titrated endogenous EF in an EwS cell line and linked phenotypic states to distinct EF thresholds. Strikingly, modest EF depletion promoted a pro-metastatic phenotype, that diminished upon near-complete EF loss. Nascent RNA sequencing revealed distinct gene clusters with heterogenous response patterns to varying EF dosage. Target genes most sensitive to subtle EF depletion contained GGAA microsatellites in EF-bound enhancers. Furthermore, we identified Krüppel-like zinc-finger transcription factors associated with EF-repressed EMP genes. Transient EF depletion followed by rapid restoration to simulate oncoprotein fluctuations identified persistently dysregulated genes associated with poor prognosis. This study underscores the therapeutic challenge of insufficient EF inhibition and provides a foundation for exploiting oncoprotein dynamics to uncover therapeutic vulnerabilities in fusion-driven cancers. Beyond EwS, our results underscore the broader impact of oncoprotein dosage dynamics in cancers with otherwise quiet genomes. SIGNIFICANCE We report EwS as a paradigm for the importance of oncogene fluctuations in tumor cell plasticity and disease progression. Effective therapeutic strategies must ensure complete EF depletion to prevent inadvertent metastasis.
Nuclear PKM2 binds pre-mRNA at folded G-quadruplexes and reveals their gene regulatory role.
In Molecular Cell on 3 October 2024 by Anastasakis, D. G., Apostolidi, M., et al.
Nuclear localization of the metabolic enzyme PKM2 is widely observed in various cancer types. We identify nuclear PKM2 as a non-canonical RNA-binding protein (RBP) that specifically interacts with folded RNA G-quadruplex (rG4) structures in precursor mRNAs (pre-mRNAs). PKM2 occupancy at rG4s prevents the binding of repressive RBPs, such as HNRNPF, and promotes the expression of rG4-containing pre-mRNAs (the "rG4ome"). We observe an upregulation of the rG4ome during epithelial-to-mesenchymal transition and a negative correlation of rG4 abundance with patient survival in different cancer types. By preventing the nuclear accumulation of PKM2, we could repress the rG4ome in triple-negative breast cancer cells and reduce migration and invasion of cancer cells in vitro and in xenograft mouse models. Our data suggest that the balance of folded and unfolded rG4s controlled by RBPs impacts gene expression during tumor progression.
Published by Elsevier Inc.
In Nature on 1 March 2023 by Wang, H., Fan, Z., et al.
Fig.3.A
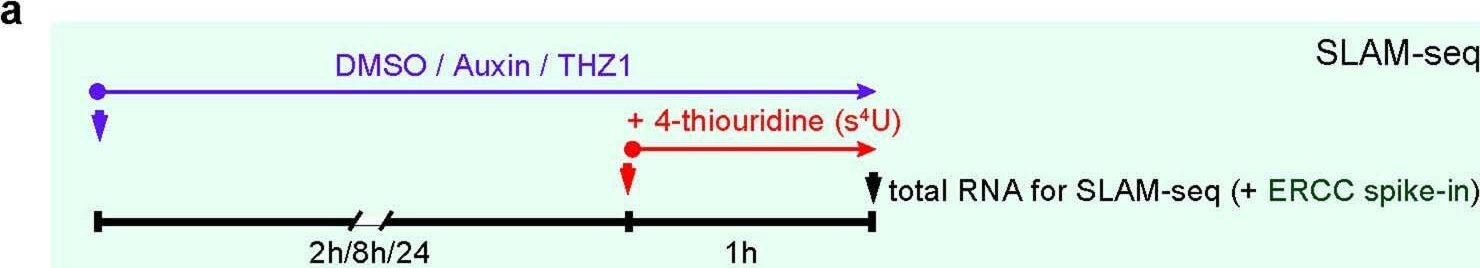
Collected and cropped from Nature by CiteAb, provided under a CC-BY license
Image 1 of 2
In Nature on 1 March 2023 by Wang, H., Fan, Z., et al.
Fig.10.A
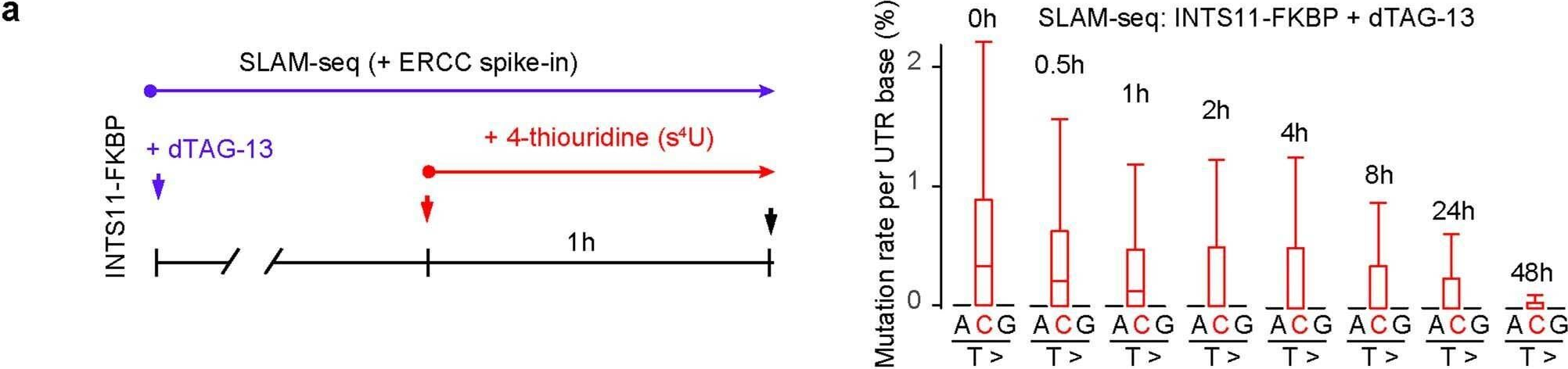
Collected and cropped from Nature by CiteAb, provided under a CC-BY license
Image 1 of 2
